Who am I?
Hi my name is Jan Heuschele and I am a researcher at the University of Oslo. In my spare time, I communicate research through visual abstracts and illustrations for grant applications, presentations, and articles. I can also help to plan and produce scientific outreach activities for your research project that goes beyond the conventional twitter feeds and Facebook posts. I have already produced children books, interactive websites, and public science installations.

The world of Hopfs
The world of Hopfs is a children's book which introduces the reader to the simple requirements of evolution: variation, heritability, and selection. The kids will discover that there are some differences between the individuals, and children inherit some of those from their parents. And sometimes the environment determines who can survive and reproduce. By now, The World of Hopfs has been translated into ten different languages. You can download most of them on the project homepage , and you can also order a paperback version there.

Earth observation brochure
Together with the National Oceanography Centre, we developed an explanatory leaflet that shows how earth observation works and how it can benefit society. The backside features a recent satellite image captured from the ESA satellites, sourced from Sentinel Hub. You can make your own personlized brochure using a Shiny app, or you can download ready made pdfs with images from beautiful places.
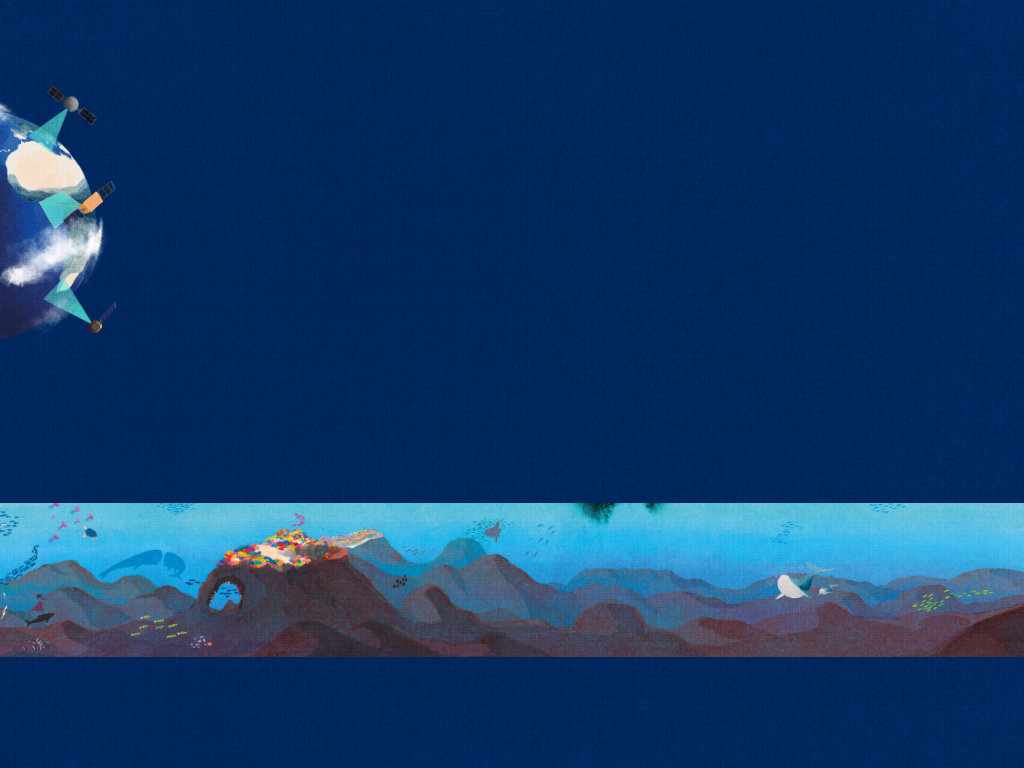

Have you seen the fish?
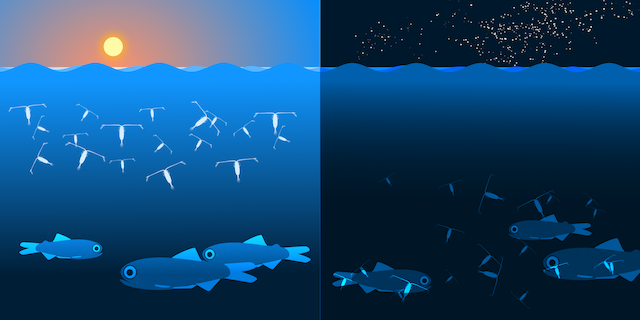
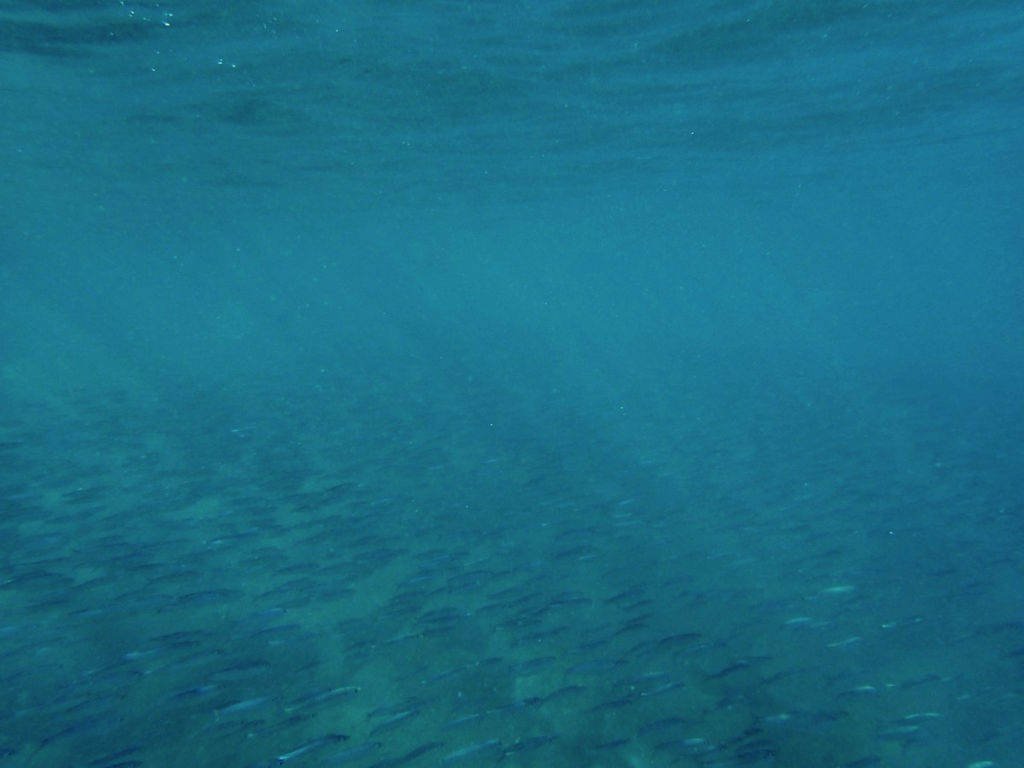
or "Har du set fisken?" in Danish, is a small picture book for children about a little fish. It was made for the research project BONUS INSPIRE to tell children about the fabulous life below the sea. In the BONUS Inspire project, researchers from nine different European countries have studied for four years some of the processes that lead to changes in fish stocks in the Baltic Sea. The goal of the book was to introduce the young reader to the complex mechanisms that lead to the different distribution of species in the ocean. You can download it here (9MB).

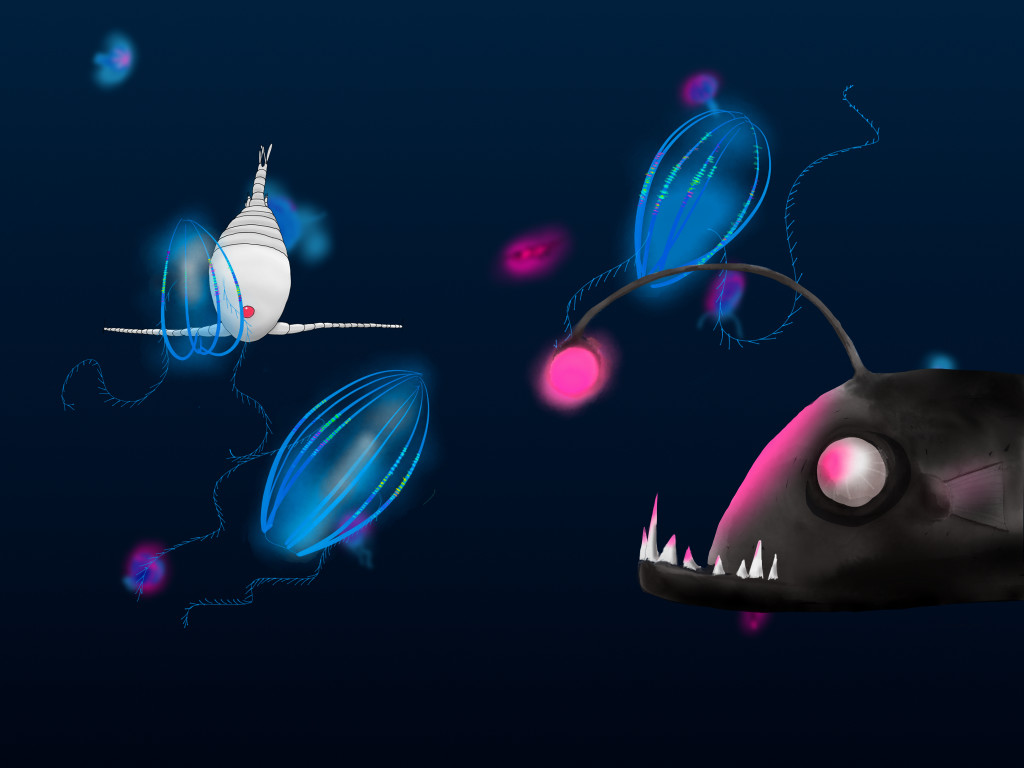
Otto the copepod
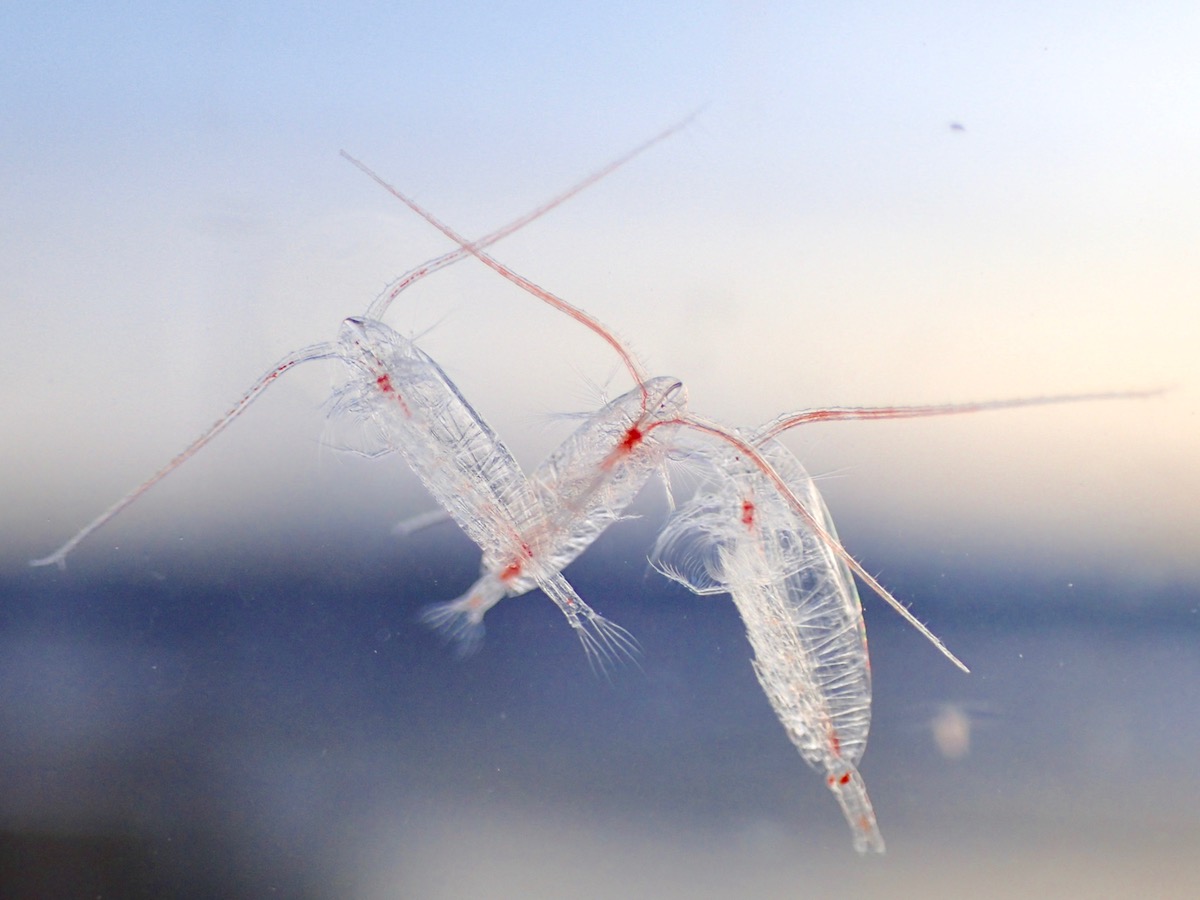
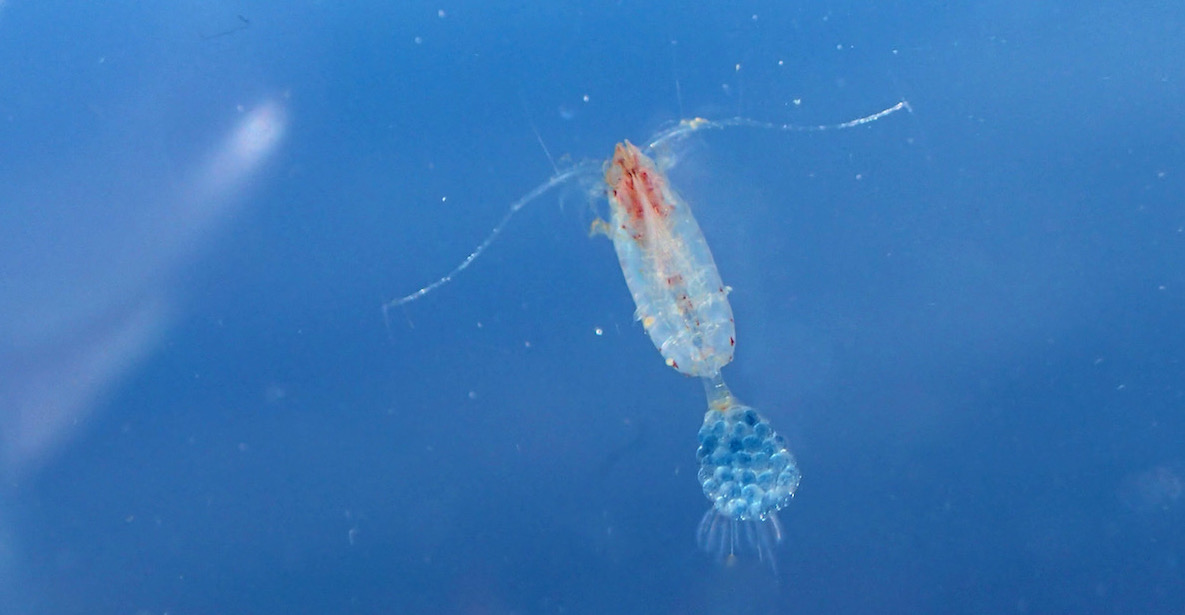
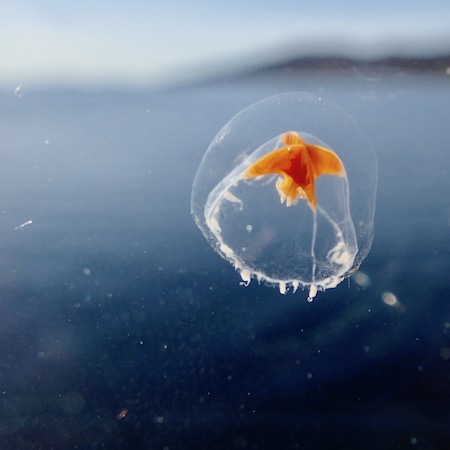
Otto the Copepod, is an interactive story about a copepod searching for its friends. On his quest, he meets all kinds of organisms. I did the project for the Centre for Ocean Life at DTU Aqua. The idea was to introduce kids and their parents to the major algae and animal groups in the open ocean. The researcher at the center also contributed by adding information about their current work at the Centre for Ocean Life. You can read it here: Otto the copepod.

Research interests
A complete and updated list of my articles can be found at ResearchGate. My research so far focused on a) the influence of anthropogenic environmental change on sexual selection in fish, b) the reproduction and development in marine zooplankton, and c) the effects of multiple stressors and parasites on aquatic invertebrates

Contact
If you have any questions or ideas on collaborations I would be delighted if you drop me a message here: jan{@}heuschele.com .
Privacy policy
I use Google Analytics on heuschele.com and the subpages “The world of Hopfs” and “Otto the copepod”. Google Analytics records data about your location, from which website you came from and for how long you stayed. It also saves what device you’re using, and a bit more. This information allows me to understand better who comes to my site and what they read. Google keeps this info for two years. Occasionally, I will compile aggregate statistics for myself about the number of visitors from different parts of the world this site receives. All of my activity falls within the bounds of the Google Analytics ToS.